Prerequisites
CRISPR: gRNA DESIGN AND VECTOR CREATION
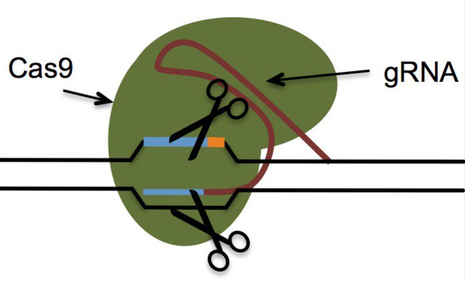
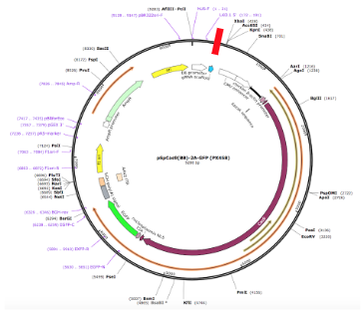
In a px458 plasmid, a gRNA was designed for two isoforms of CXCR4 CXCR4.1 at caccTCCACAGGCTATCGGGGTAA and CXCR4.2 at caccATCTTTGCCGACGTCAGCCA. Then, since the gRNA came in oligonucleotides (oligos) before the gRNA was inserted into the plasmid, the oligos were annealed together and phosphorylated on the 5’ end to prepare the gRNA for ligation [21]. The Zhang laboratory digestion and ligation reaction was used to anneal and phosphorylate the oligos. The digestion reaction exposed the sticky ends of the oligos and the ligation reaction both annealed and put phosphides on the oligos using 10µL for the final volume instead of the recommended 20µL [22]. These oligos had, by this point, been transformed to gRNA that is to be inserted into the plasmid at the BBS1 cut site at 245 and 267 bp. One set of cells had an insert, plasmid and the BBS1 cut site while the empty vector had just the plasmid and BBS1 cut site so there would be something to compare results to.
BACTERIAL TRANSFORMATION AND INSERT VERIFICATION
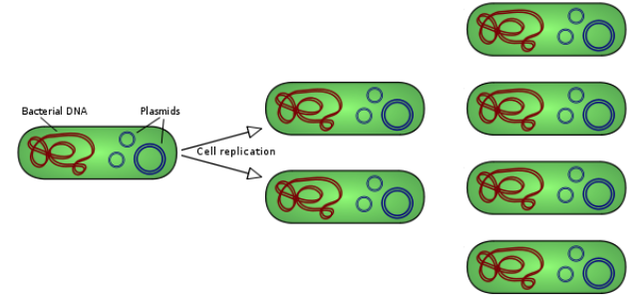
CXCR4.1 and CXCR4.2 vectors were transformed into dHα competent bacterial cells using a Transformation Protocol from Thermo Fisher [23]. Cells were then pipetted into plates that had Lysogeny broth (LB), ampicillin (amp), and agar, making sure there was 1mL of LB to 1µL of amp and 100µL of transformed cells. These cells were stored in a 37˚C bacterial incubator for about 16 hours. Then, colonies were chosen to grow in liquid LB and amp (with the same ratios as stated before) for 16 hours at 37˚C again. These colonies were shaken at 225 rpm in the incubator. With these colonies, a miniprep from Qiagen was used to isolate DNA from the bacterial plasmid. A restriction digest was run using isolated DNA with 0.5µL of BBS1, 0.5µL of Agel, 6µL of buffer O, 1µL of plasmid, and 2µL of water to amount to a final volume of 10µL. The restriction digest ran at 37˚C for 1 hour.
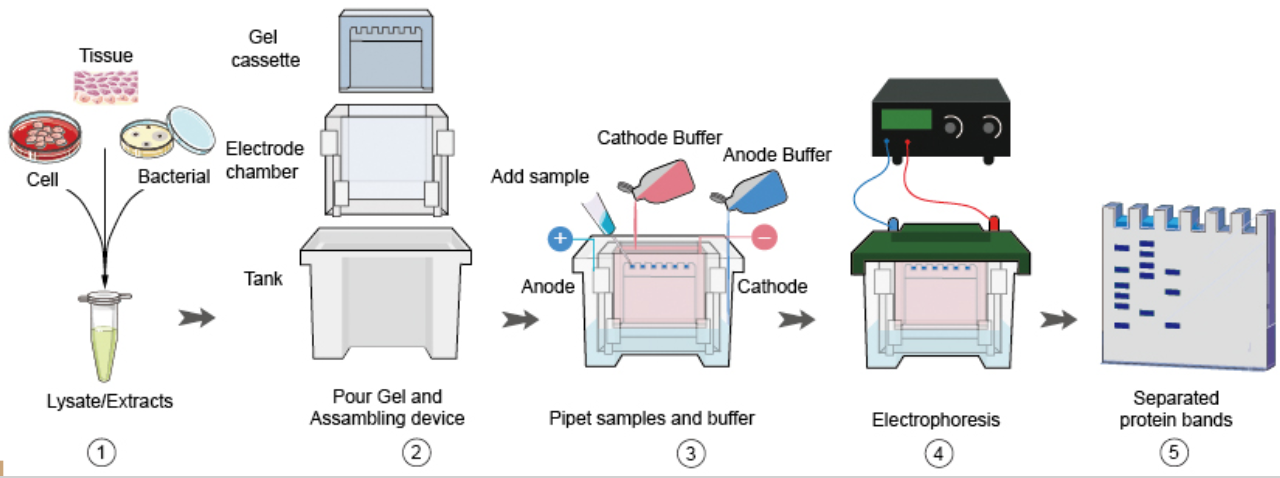
Samples were run on a 1% agarose gel to determine results.
Samples were run on a 1% agarose gel to determine results.
PREPARING FOR TRANSFECTION
Cultures were grown in 100mL of liquid LB and a midi prep from Omega Bio-tek was performed in order to isolate DNA from the cultures. On the same day, a 6-well dish with 2.0104 cells 2 days prior to transfection was prepared with 4T1 cancer cells.
TRANSFECTION
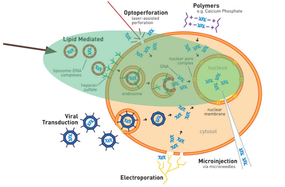
The CXCR4.1 and CXCR4.2 plasmids were transfected into the 4T1 cancer cells using the Lipofectamine 3000 Transfection Reagent protocol from Thermo Fisher Scientific, using serum free RPMI instead of optimum [23].
TRANSFECTION VERIFICATION
In order to verify the plasmid was taken in by the 4T1 BC cells, 4T1 cells were plated into culture media. To see what cells took in the plasmid, the cells were observed under a fluorescent microscope to see which cells were glowing (the plasmid had a gene that codes for the green fluorescent protein (GFP) that glows).
KNOCKOUT VERIFICATION
To see if the CXCR4 protein stopped being produced, a Western Blot was performed with the 4T1 cells that had the CXCR4.1 and CXCR4.2 plasmid inserted into them.